DCPY.DBSCANCLUST(min_samples, eps, columns)
The DBSCAN (Density Based Spatial Clustering of Applications with Noise) clustering algorithm views clusters as high-density areas separated by low density areas, when finding core samples and expanding clusters from them. Euclidean distance is used as a measure when calculating distance between data points.
Parameters
-
min_samples – The number of data points in a neighborhood for a point to be considered as a core point, integer (default 5).
-
eps – Maximum distance between two samples for them to be considered as the same neighborhood, float (default 0.5).
-
columns – Dataset columns or custom calculations.
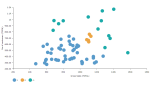
Example: DCPY.DBSCANCLUST(5, 0.5, sum([Gross Sales]), sum([No of customers])) used as a calculation for the Color field of the Scatterplot visualization.
Input data
- Numeric variables are automatically scaled to zero mean and unit variance.
- Character variables are transformed to numeric values using one-hot encoding.
- Dates are treated as character variables, so they are also one-hot encoded.
- Size of input data is not limited, but many categories in character or date variables increase rapidly the dimensionality.
- Rows that contain missing values in any of their columns are dropped.
Result
- Column of integer values starting with 1, where each number corresponds to a cluster assigned to each record (row) by the algorithm. Data points that do not belong to any cluster are considered as noise (or outliers) and are assigned to -1.
- Rows that were dropped from input data due to containing missing value have missing value instead of assigned inlier/outlier value.
Key usage points
- Automatically estimates optimum number of clusters, which can be controlled with min_samples and eps parameters.
- Not all data points are assigned to a cluster. Data points that do not belong to any cluster are considered as noise (or outliers).
- Clusters can be of any shape, but should be of similar density.
- Can find clusters completely surrounded by different clusters.
- Robust towards outliers (noise).
- Sensitivity to order of the data.
- Does not work well if clusters vary in their density.
- Not scalable with number of records and memory usage inefficiency.
- Results are very sensitive to min_samples and eps parameters.
- Suffers from 'curse of dimensionality', which may result in misleading result when the number of variables is high.
For the whole list of algorithms, see Data science built-in algorithms.